Are You Genetically Fated To Be Depressed? What I Learned From Genetic Testing and Personalized Psychiatry
TLDR
I took a stab at personalized psychiatry and found out that I have a pattern of polymorphisms that's linked to schizophrenia and treatment resistant depression. I also have a calcium channel variation that's linked to impaired cognitive ability in healthy subjects. You can learn a lot from genetic testing - but how much of the results are actionable?
Schrödinger's SNP
A few years ago I took the plunge and had some parts of my genome sequenced by a company called GenoMind. GenoMind is a company like 23andMe that leverages genetics for personalized medicine. However, GenoMind is focused on personalized psychiatry in particular.
At the time, I was worried about a Schrödinger's cat kind of situation.
What if I learned that I'm genetically predisposed to bipolar disorder? Would that make me read into symptoms and starting feeling more bipolar?
The sense that your problems can be (partially) attributed to genes might lead to fatalistic thinking.
Wearing a white coat that you believe belongs to a doctor sharply increases attention. But if you wear the same white coat believing it belongs to a painter, you will show no such improvement. Your expectations about yourself affect your performance.
Genetic testing is not cut-and-dry and my particular fears were unfounded. Genes and their regulation are extraordinarily complicated. There's no "bipolar gene" or "schizophrenia gene." Even if the function of every gene was mapped and linked to disease states, there's also still the effect of environment.
Why I Sought Genetic Testing
When the GenoMind test was ordered, I was depressed and my response to medications was lackluster. My psychiatrist suggested the idea as a kind hail mary to see if it might shed light on what medications we should try next. Also, the assay is covered by insurance.
In this post, I’m going to share the report I received from GenoMind with you. I’ll also discuss what kind of information you can glean from genetic testing. Some people think that in the future all medicine will be personalized medicine: sequencing will be as routine as prescribing drugs.
How Genetic Testing Works
"Genetic testing" brings to mind paternity conflicts. What I'm talking about is genetic testing to identify genetic variations that might provide insight into your health (or lack thereof).
You provide a genetic sample (I spit into a tube) and send it off to the lab. The lab sequences a small subset of your genome. Sequencing your entire genome would be very expensive and probably overkill. Next, the lab sends you a report highlighting a few SNPs or single nucleotide polymorphisms.
What is a SNP?
SNPs (pronounced "snips"), aka single nucleotide polymorphisms, are common types of genetic variations. The variation must be relatively common in the general population (> 1%). All SNPs are mutations but not all mutations are SNPs.
Each SNP represents a single nucleotide. For example, a SNP may replace the nucleotide adenine with the nucleotide guanine (A => G). In this case, the SNP is a missense mutation or a change in one DNA base pair that results in the substitution of one amino acid for another.
What Can You Learn From Genetic Testing?
You could learn that you have an increased risk of a specific disease, and take proactive measures to decrease your risk. Or you might discover what medicines you’re likely to have an adverse reaction to and avoid them. The hope is that you'll gain some actionable insights into your health.
Personalized medicine and pharmacogenomics are in vogue.
Simply put, pharmacogenomics is the study of how genes affect a person's response to drugs. As the word suggests, the discipline is a hybrid between pharmacology (science of drugs) and genomics (the study of genes).
Opening Pandora’s Box
Bioethics
What would you do if you knew you were going to die in 5 years?
There are many pitfalls to genetic testing. If you discovered that you had the Huntingtin protein which confers a 50% risk of dementia at age 40 on average16 - would you really want to know? Not all results are actionable.
Genetic testing raises many other questions in medical ethics, too. Should relatives of someone with a positive predictive genetic test be notified of the results and risks? What if they don't want to know?
Another pitfall stems from the fact that the interpretation of genetic testing results can be a lot like reading tea leaves. Only about 15% of human genes can be pegged to some sort of function, so it’s pretty clear that we have a lot to learn about genetics. Moreover, many associations between SNPs and disease are not as cut-and-dry as you might think.
BRCA Mutations
Consider BRCA mutations, which are extremely well-characterized.
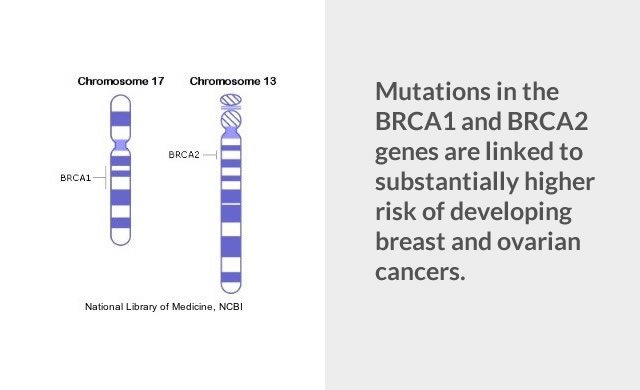
A BRCA mutation is a mutation in either of the BRCA1 and BRCA2 genes. These are tumor suppressor genes. Hundreds of mutations have been identified in these genes, but only some produce hereditary breast-ovarian cancer syndrome.
Only 5-10% of breast cancer cases in women are attributed to harmful BRCA1 and BRCA2 mutations. Having a high-risk mutation does not guarantee the development of any cancer or prove that any cancer was caused by the mutation. Yet people choose to get a mastectomy (whole breast removal) as a prophylactic measure against cancer after discovering that they have a high-risk BRCA mutation. Is this a prudent move? They might never get cancer.
Other Layers of Complexity: Alternative Splicing
Another layer of complexity arises from alternative splicing. One gene can encode many proteins; a particular exon can be included or excluded from the processed mRNA.
Alternative splicing allows the human genome to direct the synthesis of many more proteins than would be expected from its 20,000 protein-coding genes.
A Quick Refresher on Notation In Genetics
Consider the statement:
The A-allele at rs1006737 within CACNA1C ...
What does this mean?
rs1006737 - The rs number stands for Reference SNP cluster ID. It is an accession number used by researchers and databases to refer to specific SNPs.
CACNA1C - The gene that's affected by the SNP.
A-allele - an allele is one of two or more alternative forms of a gene that arise by mutation and are found at the same place on a chromosome. The "A" in "A-allele" refers to the nucleotide that's been changed (in this case, adenine). The most common genotype is G/G, i.e., having guanine at this position on both copies of the gene. G/A and A/A are the genotypes that predispose to disease.
GenoMind Results
 GenoMind sequenced part of my genome.
GenoMind sequenced part of my genome.
I had potentially deleterious genetic variations in the following:
- SLC6A4 encoding the serotonin transporter (S/S genotype)
- 5-HT2C - encodes the 5-HT2C serotonin receptor (C/C genotype)
- CACNA1C - encodes a subunit of L-type calcium channels (G/A genotype)
Of particular note: my calcium channel genotype (CACNA1C G/A) is linked to mental illness and slightly diminished working memory.
SLC6A4 (Serotonin Transporter)
The SLC6A4 gene encodes none other than the serotonin transporter (SERT).
SERT is a membrane protein that pumps serotonin back into presynaptic neurons. Heightened activity of this pump depletes serotonin; reduced pump activity prolongs the action of serotonin. SSRIs (selective serotonin reuptake inhibitors) improve mood by blocking the activity of the serotonin transporter.
SLC6A4 and Disease
I have the S/S variant of SLC6A4. This variant is actually not a SNP - it refers to the variable number of tandem repeats (VNTRs). Researchers commonly describe two variations in humans: A short ("S") and a long ("L"), which can actually be subdivided further.
The SLC6A4 S/S variation is associated with:
- Poor response to antidepressants and treatment-resistant depression1
- Increased risk of developing serotonin syndrome when treated with serotonergic drugs
- Impaired resilience to stressful life events
A little over 1/5 of caucasians have this genotype, but it’s the most common genotype in Asians and American-Indians.
Here’s the distribution of genotypes by ethnicity:

5-HTTLPR is shorthand for the "serotonin-transporter-linked polymorphic region." It is a degenerate repeat polymorphic region in SLC6A4
Since its identification in the 90's, 5-HTTLPR has been extensively investigated. There's been a big effort to link variations to neuropsychiatric disorders.
The SLC6A4 short ("S") variation decreases the activity of the serotonin transporter. Hence, it's no surprise that the S variation is linked to poor response to serotonergic drugs (e.g., SSRIs) and an increased risk of serotonin syndrome. Coversely, individuals with the long ("L") variation or the Stin2 12/12 genotype were more likely to achieve remission from depression with citalopram treatment.
Of note, it is possible that relatively mild 5-HT syndrome occurrence may contribute to early discontinuation of SRIs and side effects during SRI treatment that are strongly associated with the SS genotype and S allele of the SERT variant [18].
Meta-analyses of SLC6A4 variants have found many associations:
- Bipolar affective (5HTTLPR, S allele) 7
- Autism (5HTTLPR, S allele) 8
- Substance dependence (5HTTLPR, S allele)
- Schizophrenia (Stin2) (~12 studies, >2000 cases and >2000 controls) 9
- Depression and anxiety (~1000 cases + ~1000 controls) 10
- Obsessive-compulsive disorder (5HTTLPR L allele in family-based and case-control studies involving children and Caucasian samples, some including rs25531 and rs25532 and haplotype analyses) 11
- ADHD is linked to the 5HTTLPR L allele only, not Stin2 or rs3813034 in a meta-analysis of 20 studies 12
In brief, having the SS genotype is far from ideal.
5-HT2C
5-HT2C is a subtype of the serotonin receptor. I had the C/C genotype (accession number: rs1414334), which is linked to increased adverse effects with antipsychotic treatment.
Notable SNPs in 5-HT2C
Interesting SNPs affecting the 5-HT2C serotonin receptor:
- rs1414334 (risk allele: C), rs3813928, rs498177, rs3813928, rs498207, rs521018 - associated with the metabolic syndrome in patients treated with antipsychotics
- rs2192372, rs2428707, rs4272555 (risk allele: T) - linked to suicidal behavior
- rs518147 - protective effect against weight-gain in antipsychotic treated patients
- rs3813929 (risk allele: T) - obesity, increased incidence of depression, adverse effects with antipsychotic treatment
- rs6318 (risk allele: C) - increased cortisol, risk of cardiac events, antipsychotic-induced weight gain
5-HT2C and Novelty Seeking Behavior
Belmaker RH's group reported a link between 5-HT2C serotonin receptor gene polymorphisms and the human personality of reward dependence 12.
The D4 dopamine receptor polymorphism (D4DR) explains a small percent of the variance for the trait of novelty seeking. Therefore the authors investigated two other coding region polymorphisms. The authors looked at a glycine => serine substitution in the dopamine D3 receptor (D3DR) and a cysteine => serine substitution in the 5-HT2C serotonin receptor. The presence of the less common 5-HT2C serine polymorphism reportedly reduces reward dependence/novelty seeking behavior.
Three-way analysis of variance (TPQ score grouped by D4DR, D3DR and 5-HT2C) demonstrated that reward dependence and persistence scores were significantly reduced by the presence of the less common 5-HT2Cser polymorphism.
5-HT2C (HTR2C) serotonin receptor gene polymorphism associated with the human personality trait of reward dependence: interaction with dopamine D4 receptor (D4DR) and dopamine D3 receptor (D3DR) polymorphisms.
5-HT2C and Adverse Effects of Antipsychotics
Zhi Jun Zhang and colleagues investigated whether 5-HT2C receptor polymorphisms affect patient's response to antipsychotic treatment. The adverse effects of antipsychotics - particularly weight gain and tardive dyskinesia - are intensified in individuals polymorphisms affecting 5-HT2C.
Calcium Channels (CACNA1C)
CACNA1C is a gene that encodes the alpha-1C subunit of the voltage-gated L-type calcium channel. This is the major constituent of this type of calcium channel in the brain. Such channels transport positively charged calcium ions into cells. Calcium channels therefore provide a key role in the cell's ability to propagate signals.
L-type calcium channels also play an important role in dendritic development, neuronal survival, synaptic plasticity and learning/memory 4
My Results
I had the G/A polymorphism (accession number: rs1006737), which is common variation linked to altered calcium channels. Alterations in calcium channels can affect brain cell excitability and may contribute to mood instability (e.g., bipolar disorder).
Examples of neuronal calcium channels include the NMDA-type glutamate receptor, which underpins long-term potentiation. This receptor is the neuronal target of many nootropics and neurotoxins alike.
Genetic variations in CACNA1C affects brain circuitries related to mental illness. 2 Specifically, CACNA1C gene polymorphisms are linked to bipolar disorder3, schizophrenia, and unipolar depression.
The CACNA1C A-Allele Impairs Cognitive Performance in Schizophrenia and Healthy Controls

Comparisons of performance on the seven neurocognitive domains between A-allele carriers (white bars) and G-allele homozygotes (black bars) in patients with schizophrenia (a) and control subjects (b).2
CACNA1C and Disease
The A-allele at rs1006737 within CACNA1C is associated with:
- Global grey matter volume in a sample of 77 adults17
- In a different study3, global grey matter volume was also associated with rs1006737, with volume decreasing with the total number of A alleles
- Spatial working memory performance in a knockdown mouse model18
- Impaired working memory in schizophrenia patients19
- Decreased attention, working memory, and verbal fluency in healthy adults5
- Increased amygdala volume20
Are You Genetically Fated To Be Depressed?
I think the answer is a resounding "no."
References
-
Taylor MJ, Sen S, Bhagwagar Z. Antidepressant response and the serotonin transporter gene-linked polymorphic region. Biol Psychiatry. 2010;68(6):536-43. ↩
-
Bigos KL, Mattay VS, Callicott JH, et al. Genetic variation in CACNA1C affects brain circuitries related to mental illness. Arch Gen Psychiatry. 2010;67(9):939-45. ↩
-
Franke B, Vasquez AA, Veltman JA, Brunner HG, Rijpkema M, Fernández G. Genetic variation in CACNA1C, a gene associated with bipolar disorder, influences brainstem rather than gray matter volume in healthy individuals. Biol Psychiatry. 2010;68(6):586-8. ↩
-
Bhat S. et al. CACNA1C (Ca(v)1.2) in the pathophysiology of psychiatric disease. Prog Neurobiol in press. ↩
-
Zhang Q. J. et al. The effects of CACNA1C gene polymorphism on spatial working memory in both healthy controls and patients with schizophrenia or bipolar disorder. Neuropsychopharmacology 37, 677–684 (2012). [PMC free article] [PubMed] [Ref list] ↩
-
Thimm M. et al. Effects of a CACNA1C genotype on attention networks in healthy individuals. Psychol Med 41, 1551–1561 (2011). [PubMed] [Ref list] ↩
-
Cho HJ, Meira-Lima I, Cordeiro Q, Michelon L, Sham P, Vallada H, Collier DA. Population-based and family-based studies on the serotonin transporter gene polymorphisms and bipolar disorder: a systematic review and meta-analysis. Mol Psychiatry. 2005;10:771–781 ↩
-
Kistner-Griffin E, Brune CW, Davis LK, Sutcliffe JS, Cox NJ, Cook EH., Jr Parent-of-origin effects of the serotonin transporter gene associated with autism. Am J Med Genet B Neuropsychiatr Genet. 2010 ↩
-
Fan JB, Sklar P. Meta-analysis reveals association between serotonin transporter gene STin2 VNTR polymorphism and schizophrenia. Mol Psychiatry. 2005;10:928–938. 891. ↩
-
Wray NR, James MR, Gordon SD, Dumenil T, Ryan L, Coventry WL, Statham DJ, Pergadia ML, Madden PA, Heath AC, et al. Accurate, Large-Scale Genotyping of 5HTTLPR and Flanking Single Nucleotide Polymorphisms in an Association Study of Depression, Anxiety, and Personality Measures. Biol Psychiatry. 2009;66:468–476. ↩
-
Bloch MH, Landeros-Weisenberger A, Sen S, Dombrowski P, Kelmendi B, Coric V, Pittenger C, Leckman JF. Association of the serotonin transporter polymorphism and obsessive-compulsive disorder: systematic review. Am J Med Genet B Neuropsychiatr Genet. 2008;147B:850–858. ↩
-
https://ghr.nlm.nih.gov/gene/CACNA1C#conditions ↩
-
https://www.ncbi.nlm.nih.gov/pubmed/20638048 ↩
-
Myers RH. Huntington's disease genetics. NeuroRx. 2004;1(2):255-62. ↩
-
Kempton MJ, Ruberto G, Vassos E, Tatarelli R, Girardi P, Collier D, Frangou S (2009): Effects of the CACNA1C risk allele for bipolar disorder on cerebral gray matter volume in healthy individuals. Am J Psychiatry 166:1413–1414. ↩
-
Moosmang S, Haider N, Klugbauer N, Adelsberger H, Langwieser N, Muller J, et al. (2005): Role of hippocampal Cav1.2 Ca2 channels in NMDA receptor-independent synaptic plasticity and spatial memory. J Neurosci 25:9883–9892. ↩
-
GreenEK,GrozevaD,JonesI,JonesL,KirovG,CaesarS,etal.(2009):The bipolar disorder risk allele at CACNA1C also confers risk of recurrent major depression and of schizophrenia [published online ahead of print July 21]. Mol Psychiatry. ↩
-
Lancaster TM, Foley S, Tansey KE, Linden DE, Caseras X. CACNA1C risk variant is associated with increased amygdala volume. Eur Arch Psychiatry Clin Neurosci. 2016;266(3):269-75. ↩

